Chapter 9: Bayesian Predictive Analysis
Chapter09.RmdSection 9.1: From Bayesian Posterior to Bayesian Predictive Inference
Example 9.1: Road Safety Data - Single Predictions
We load the data and extract the observations for the senior people in Linz. We then plot the pdf and cdf for the predictive distribution which corresponds under the flat prior to
data("accidents", package = "BayesianLearningCode")
y <- accidents[, "seniors_accidents"]
(aN <- sum(y) + 1)
#> [1] 1009
(bN <- length(y))
#> [1] 192
mu <- aN / bN
yf <- 0:20
plot(yf, dnbinom(yf, size = aN, mu = mu),
type = "h", xlab = bquote(y[f]), ylab = "", lwd = 2)
plot(yf, pnbinom(yf, size = aN, mu = mu),
type = "h", xlab = bquote(y[f]), ylab = "", lwd = 2)
probs <- c(0.025, 0.975)
abline(h = probs, lty = 3)
mtext(probs, side = 2, at = probs, adj = c(0, 1), cex = .8, col = "dimgrey")
Example 9.2: Exchange Rate Data
We load the data and then plot the pdf and cdf for the predictive distribution which corresponds under the improper prior to
library("BayesianLearningCode")
data("exrates", package = "stochvol")
y <- 100 * diff(log(exrates$USD / exrates$CHF))
ybar <- mean(y)
N <- length(y)
b0 <- 0
N0 <- 0
c0 <- -1/2
C0 <- 0
BN <- 1 / (N + N0)
bN <- BN * (N * ybar + N0 * b0)
cN <- c0 + N/2
CN <- C0 + .5 * sum((y - ybar)^2) + N0 * N / (2 * (N0 + N)) * (b0 - ybar)^2
x <- seq(-3, 3, length.out = 200)
scale <- sqrt(CN / cN * (BN + 1))
plot(x, dstudt(x, location = bN, scale = scale, df = 2 * cN),
type = "l", xlab = bquote(y[f]), ylab = "", lwd = 1.5)
plot(x, pstudt(x, location = bN, scale = scale, df = 2 * cN),
type = "l", xlab = bquote(y[f]), ylab = "", lwd = 1.5)
probs <- c(0.025, 0.975)
abline(h = probs, lty = 3)
mtext(probs, side = 2, at = probs, adj = c(0, 1), cex = .8, col = "dimgrey")
We inspect the parameters of the Student-t distribution.
Example 9.3: Exchange Rate Data cont’d
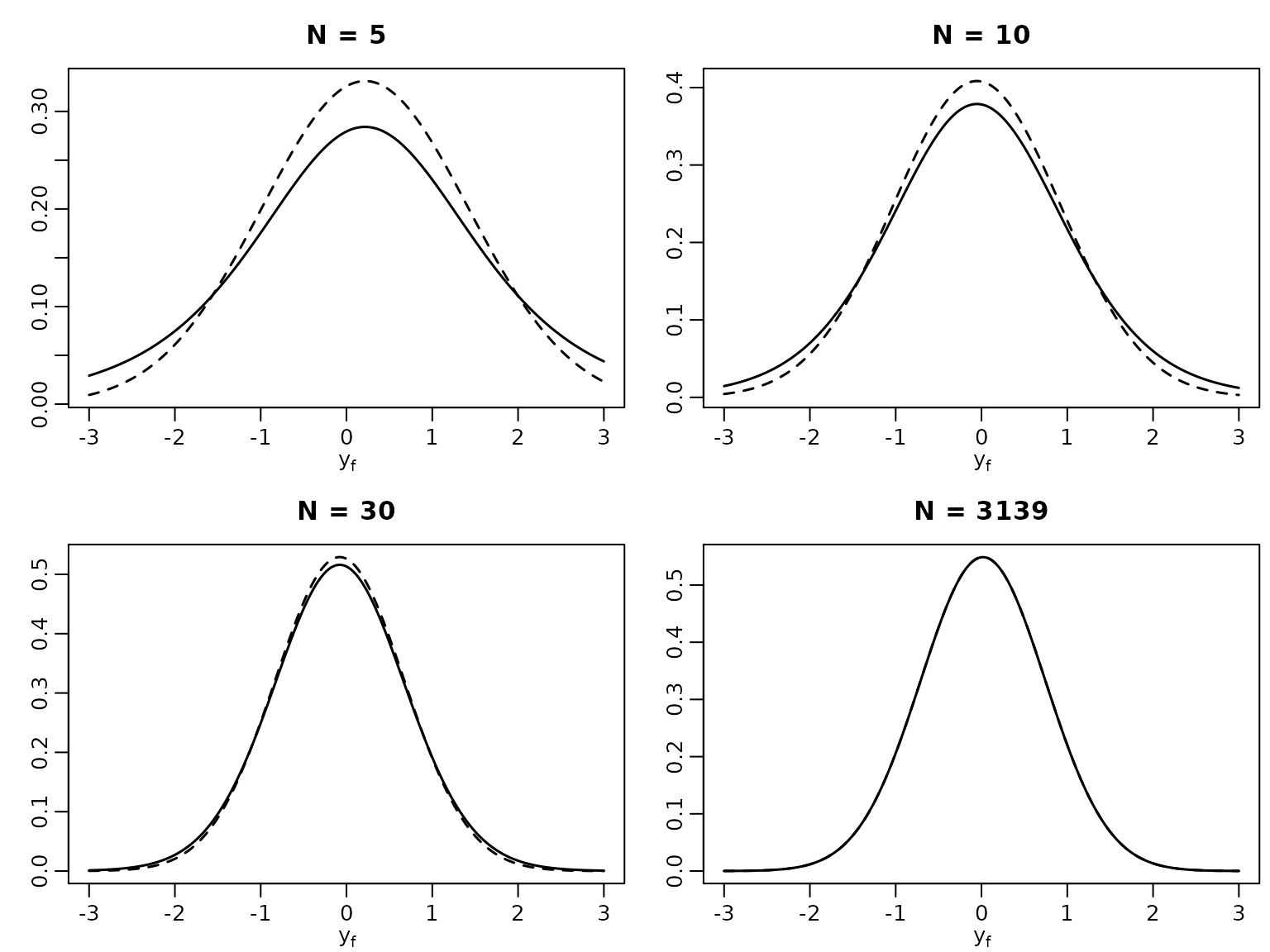
To compare with a method that ignores parameter uncertainty, we now plot the posterior predictive alongside the “classical” forecasting distribution for varying .
Ns <- c(5, 10, 30, N)
for (i in seq_along(Ns)) {
N <- Ns[i]
yshort <- y[1:N]
ybar <- mean(yshort)
BN <- 1 / (N + N0)
bN <- BN * (N * ybar + N0 * b0)
cN <- c0 + N/2
CN <- C0 + .5 * sum((yshort - ybar)^2) + N0 * N / (2 * (N0 + N)) * (b0 - ybar)^2
scale <- sqrt(CN / cN * (BN + 1))
plot(x, dnorm(x, ybar, sd(yshort)), lty = 2, type = "l", lwd = 1.5,
xlab = bquote(y[f]), ylab = "", main = paste("N =", N))
lines(x, dstudt(x, location = bN, scale = scale, df = 2 * cN), lwd = 1.5)
}
Example 9.4: Road Safety Data cont’d
We verify that a 95% predictive interval is given by [1, 9] using the cdf and compute the effective coverage.
Example 9.5: Exchange Rate Data cont’d
We determine the 95% predictive interval using the quantile function.
We now work out the effective coverage of the naive interval which ignores parameter uncertainty.
coverage <- rep(NA_real_, length(Ns))
names(coverage) <- Ns
for (i in seq_along(Ns)) {
N <- Ns[i]
yshort <- y[1:N]
quants <- qnorm(probs, mean(yshort), sd(yshort))
coverage[i] <- diff(pstudt(quants, mean(yshort), sd(yshort) * sqrt(1 + 1/N),
2 * (c0 + N/2)))
}
knitr::kable(t(round(coverage, 4)))| 5 | 10 | 30 | 3139 |
|---|---|---|---|
| 0.8519 | 0.9055 | 0.9363 | 0.9499 |
Section 9.3: A Sampling-Based Approach to Prediction
Example 9.8: Sampling-based prediction for the CHF/USD exchange rates
We proceed exactly as in Chapter 4. For the Gaussian distribution, the posterior of is inverse gamma, and we can easily generate iid draws.
For the Student- model, we can use inverse transform sampling. First, we draw uniformly from the interval spanned by 0 and the maximum of the non-normalized cumulative posterior. Then, for each draw, we find the interval of our pointwise cdf approximation of the posterior, and interpolate linearly between the interval boundaries.
# We need the nonnormalized cumulative posterior distribution
post_nonnormalized_nonvec <- function(sigma2, y, nu, log = FALSE) {
logdens <- -length(y) / 2 * log(sigma2) -
(nu + 1) / 2 * sum(log(1 + y^2 / (nu * sigma2))) - log(sigma2)
if (log) logdens else exp(logdens)
}
post_nonnormalized <- Vectorize(post_nonnormalized_nonvec, "sigma2")
# Now, we compute the pdf and cdf on a reasonably chosen grid
nu <- 7
sigma2 <- seq(0.25, 0.45, length.out = 3000)
pdf_u <- post_nonnormalized(sigma2, y = y, nu = nu)
pdf_u <- pdf_u / max(pdf_u)
cdf_u <- cumsum(pdf_u) / sum(pdf_u)
# Now we can perform inverse transpose sampling
unifdraws <- runif(ndraws, 0, cdf_u[length(cdf_u)])
leftind <- findInterval(unifdraws, cdf_u)
rightind <- leftind + 1L
distprop <- (unifdraws - cdf_u[leftind]) / (cdf_u[rightind] - cdf_u[leftind])
sigma2draws_t <- sigma2[leftind] + distprop *
(sigma2[rightind] - sigma2[leftind])Next, we simulate draws from the posterior predictive.
yf_normal <- rnorm(ndraws, 0, sqrt(sigma2draws_normal))
yf_t <- rstudt(ndraws, 0, sqrt(sigma2draws_t), 7)Now we can compute their empirical quantiles, and those of the data.
quants <- c(0.01, 0.05, 0.25, 0.4, 0.5, 0.6, 0.75, 0.95, 0.99)
q_normal <- quantile(yf_normal, quants)
q_t <- quantile(yf_t, quants)
q_y <- quantile(y, quants)
knitr::kable(round(t(cbind("Data" = q_y, "Student t" = q_t,
"Gaussian" = q_normal)), 3))| 1% | 5% | 25% | 40% | 50% | 60% | 75% | 95% | 99% | |
|---|---|---|---|---|---|---|---|---|---|
| Data | -1.687 | -1.078 | -0.416 | -0.137 | 0.005 | 0.159 | 0.435 | 1.158 | 1.864 |
| Student t | -1.721 | -1.119 | -0.421 | -0.149 | 0.004 | 0.164 | 0.432 | 1.125 | 1.741 |
| Gaussian | -1.676 | -1.185 | -0.487 | -0.177 | 0.008 | 0.185 | 0.507 | 1.211 | 1.704 |
We conclude by visualizing the data and the predictive distributions.
minmax <- ceiling(10 * max(abs(y))) / 10
grid <- seq(-minmax, minmax, length.out = 50)
hist(y, freq = FALSE, breaks = grid, border = NA,
main = "Histogram and predictive densitites")
lines(density(yf_normal, adjust = 2), col = 4, lty = 1, lwd = 1.5)
lines(density(yf_t, adjust = 2), col = 2, lty = 2, lwd = 1.5)
legend("topleft", c("Normal", "Student t"), lty = 1:2, col = c(4,2), lwd = 1.5)
ts.plot(y, main = "Time series plot and some predictive quantiles")
abline(h = q_normal, col = 4, lty = 1, lwd = 1.5)
abline(h = q_t, col = 2, lty = 2, lwd = 1.5)
legend("topleft", c("Normal", "Student t"), lty = 1:2, col = c(4,2), lwd = 1.5)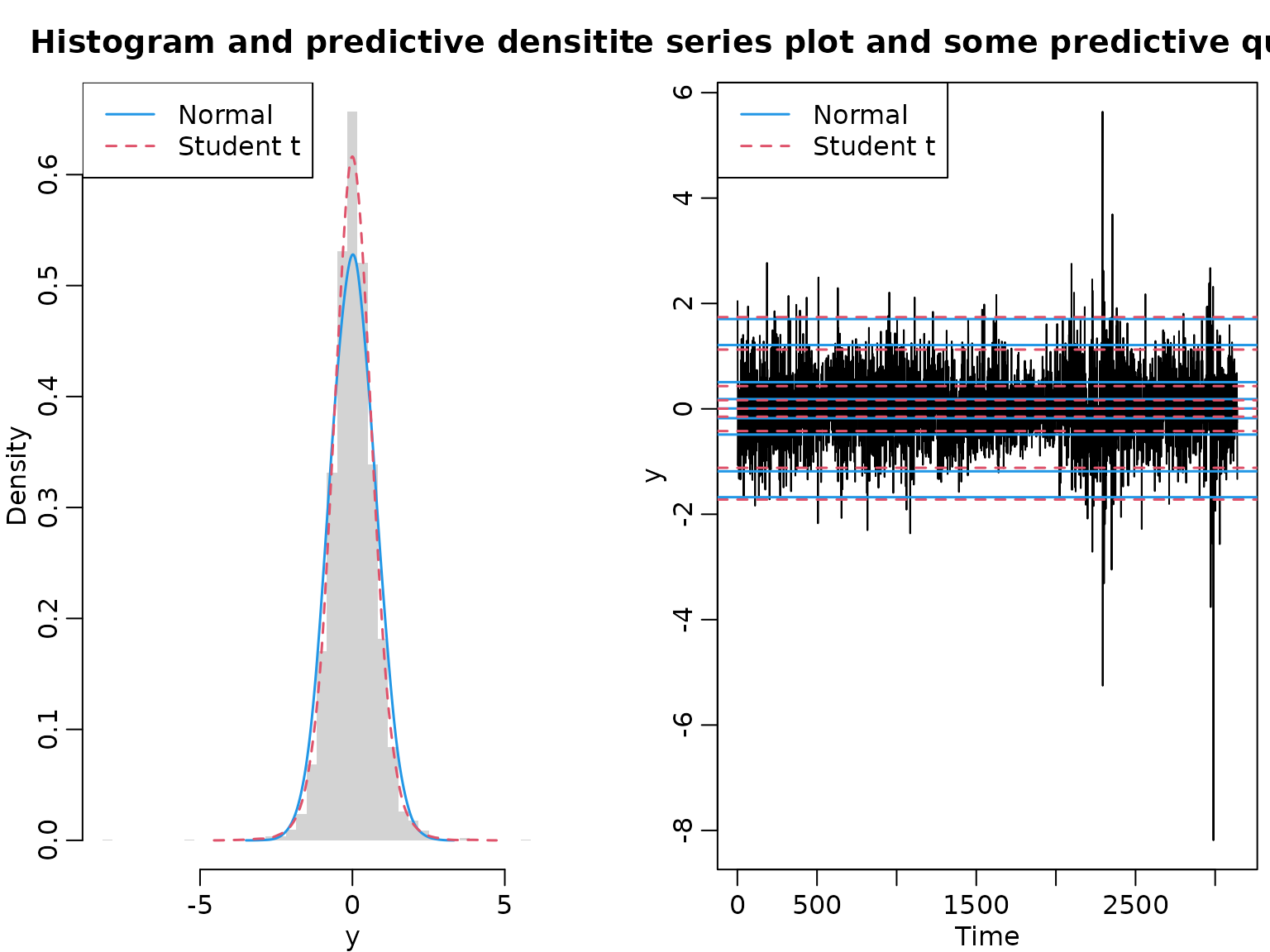
Example 9.9: Predicting yearly maxima for the road safety data
We use a sampling-based approach to obtain draws from posterior predictive by first drawing from the posterior . Then, using these draws as mean parameters for the Poisson likelihood, we draw 12 times each to obtain yearly predictions. Of these, we take the maxima.
set.seed(1)
y <- accidents[, "seniors_accidents"]
aN <- sum(y) + 1
bN <- length(y)
mus <- rgamma(ndraws, aN, bN)
yfs <- matrix(rpois(12 * ndraws, mus), ncol = 12)
Us <- apply(yfs, 1, max)Now we visualize.
plot(tab <- proportions(table(Us)), xlab = "U", ylab = "")
plot(as.table(cumsum(tab)), type = "h", xlab = "U", ylab = "")
probs <- c(0.025, 0.975)
abline(h = probs, lty = 3)
mtext(probs, side = 2, at = probs, adj = c(0, 1), cex = .8, col = "dimgrey")
Example 9.11
We illustrate the variance of the purely sampling-based estimator versus the Rao-Blackwellized version by running several experiments.
set.seed(42)
a0 <- 1
b0 <- 1
N <- 100
SN <- 42
H <- 10
M <- 100
nsim <- 1000
# compute the posterior parameters:
aN <- a0 + SN
bN <- b0 + N - SN
mu <- aN / bN
pk1 <- pk2 <- matrix(NA_real_, nsim, H + 1)
colnames(pk1) <- colnames(pk2) <- 0:10
for (i in seq_len(nsim)) {
# purely sampling-based:
thetas <- rbeta(M, aN, bN)
Sfs <- rbinom(M, H, thetas)
pk1[i, ] <- proportions(tabulate(Sfs + 1, nbins = 11)) # tabulate starts at 1
# Rao-Blackwellized:
for (k in 0:H) {
pk2[i, k + 1] <- mean(dbinom(k, H, thetas))
}
}To compute the exact probabilities, we need the density function of the beta-binomial distribution (which is not part of base R).
dbetabinom <- function(x, n, a, b, log = FALSE) {
logdens <- lchoose(n, x) + lbeta(x + a, n - x + b) - lbeta(a, b)
if (log) logdens else exp(logdens)
}Now we can easily evaluate the probabilities.
pk3 <- dbetabinom(0:H, H, aN, bN)To concluse, we visualize.
boxplot(pk1, xlab = "k", range = 0, main = "Purely sampling-based")
points(pk3, col = 3, cex = 1.5, pch = 16)
boxplot(pk2, xlab = "k", range = 0, main = "Rao-Blackwellized",
ylim = range(pk1))
points(pk3, col = 3, cex = 1.5, pch = 16)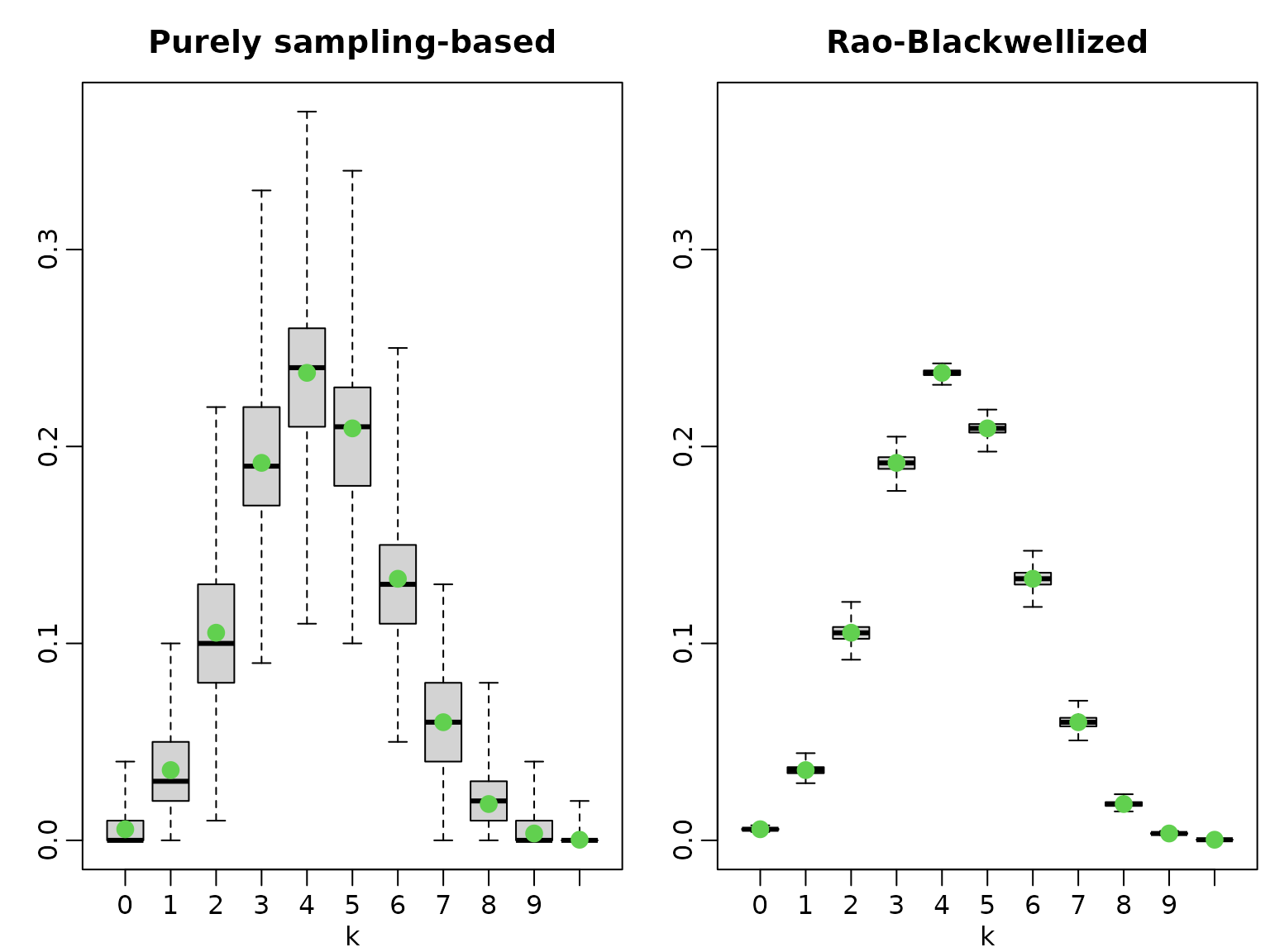
Section 9.5: Bayesian Forecasting of Time Series
Example 9.14: One-step-ahead forecasting of GDP
For creating the design matrix for an AR model, we re-use the function from Chapter 7.
ARdesignmatrix <- function(dat, p = 1) {
d <- p + 1
N <- length(dat) - p
Xy <- matrix(NA_real_, N, d)
Xy[, 1] <- 1
for (i in seq_len(p)) {
Xy[, i + 1] <- dat[(p + 1 - i) : (length(dat) - i)]
}
Xy
}We compute the one-step-ahead posterior predictive for various AR() models under the improper prior.
data(gdp)
dat <- gdp[1:which(names(gdp) == "2019-10-01")]
logret <- diff(log(dat))
means <- scales <- dfs <- rep(NA_real_, 4)
for (p in 1:4) {
y <- tail(logret, -p)
X <- ARdesignmatrix(logret, p)
N <- nrow(X)
d <- ncol(X)
BN <- solve(crossprod(X))
bN <- tcrossprod(BN, X) %*% y
SSR <- sum((y - X %*% bN)^2)
cN <- (N - d) / 2
CN <- SSR / 2
xf <- c(1, rev(tail(y, p)))
means[p] <- xf %*% bN
scales[p] <- sqrt((crossprod(xf, BN) %*% xf + 1) * CN / cN)
dfs[p] <- 2 * cN
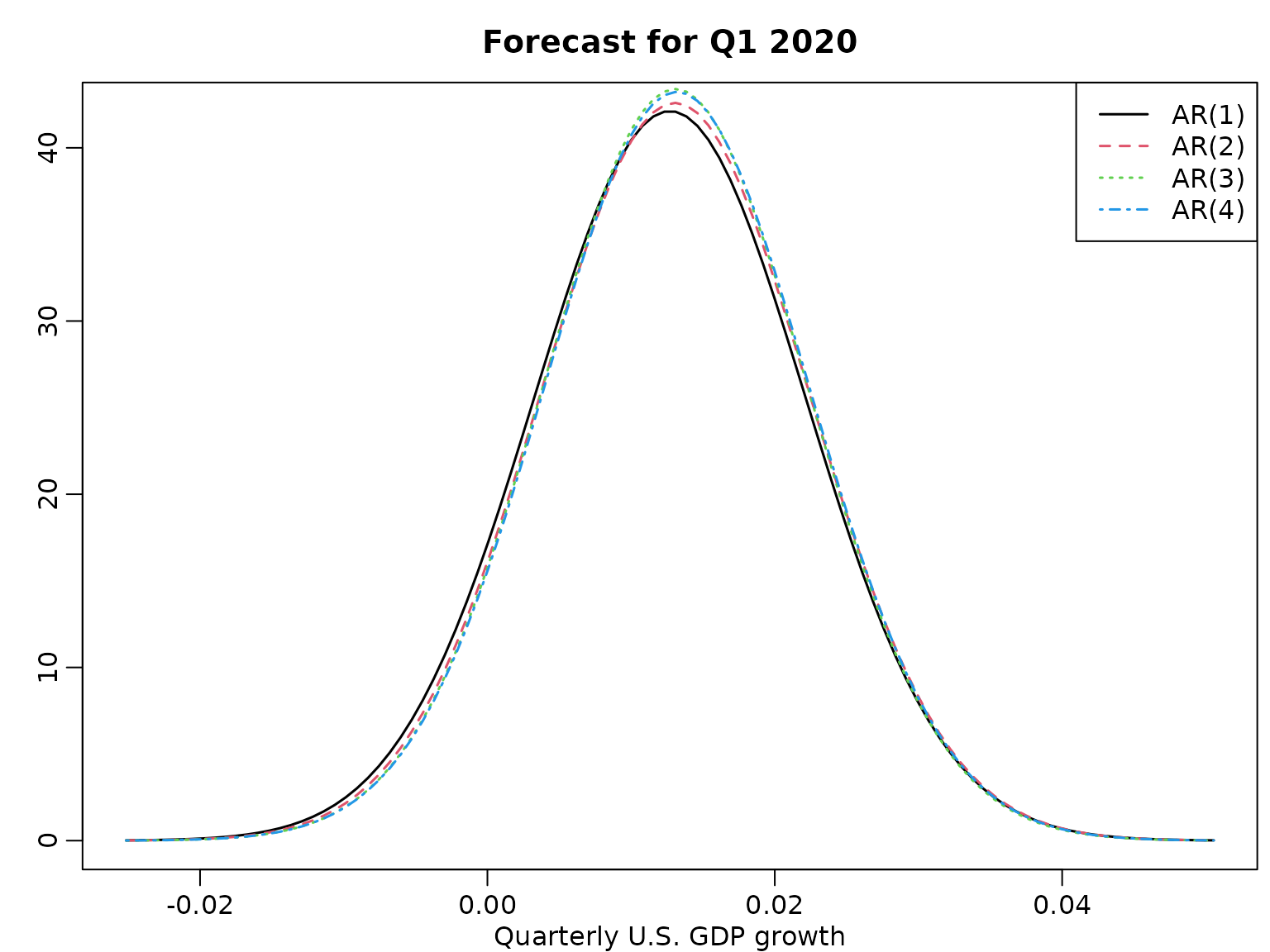
}And we visualize.
library(BayesianLearningCode)
grid <- seq(min(means - 4 * scales), max(means + 4 * scales), length.out = 100)
plot(grid, dstudt(grid, means[1], scales[1], dfs[1]), type = "l",
ylab = "", xlab = "Quarterly U.S. GDP growth", lwd = 1.5)
title("Forecast for Q1 2020")
legend("topright", paste0("AR(", 1:4, ")"), lty = 1:4, col = 1:4, lwd = 1.5)
for (p in 2:4) {
lines(grid, dstudt(grid, means[p], scales[p], dfs[p]), col = p, lty = p,
lwd = 1.5)
}
Now, we want to “sample the future” up to 12 steps ahead for .
M <- 10000
set.seed(1)
y <- tail(logret, -2)
X <- ARdesignmatrix(logret, 2)
N <- nrow(X)
d <- ncol(X)
BN <- solve(crossprod(X))
bN <- tcrossprod(BN, X) %*% y
SSR <- sum((y - X %*% bN)^2)
cN <- (N - d) / 2
CN <- SSR / 2
sigma2s <- rinvgamma(M, cN, CN)
betas <- matrix(NA_real_, M, 3)
for (i in seq_len(M)) betas[i, ] <- mvtnorm::rmvnorm(1, bN, sigma2s[i] * BN)
zetas <- betas[, 1]
phi1s <- betas[, 2]
phi2s <- betas[, 3]
sigmas <- sqrt(sigma2s)
yfs <- matrix(NA_real_, M, 8)
yfs[, 1] <- rnorm(M, zetas + phi1s * y[length(y)] +
phi2s * y[length(y) - 1], sigmas)
yfs[, 2] <- rnorm(M, zetas + phi1s * yfs[, 1] +
phi2s * y[length(y)], sigmas)
for (h in 3:8) {
yfs[, h] <- rnorm(M, zetas + phi1s * yfs[, h - 1] + phi2s * yfs[, h - 2],
sigmas)
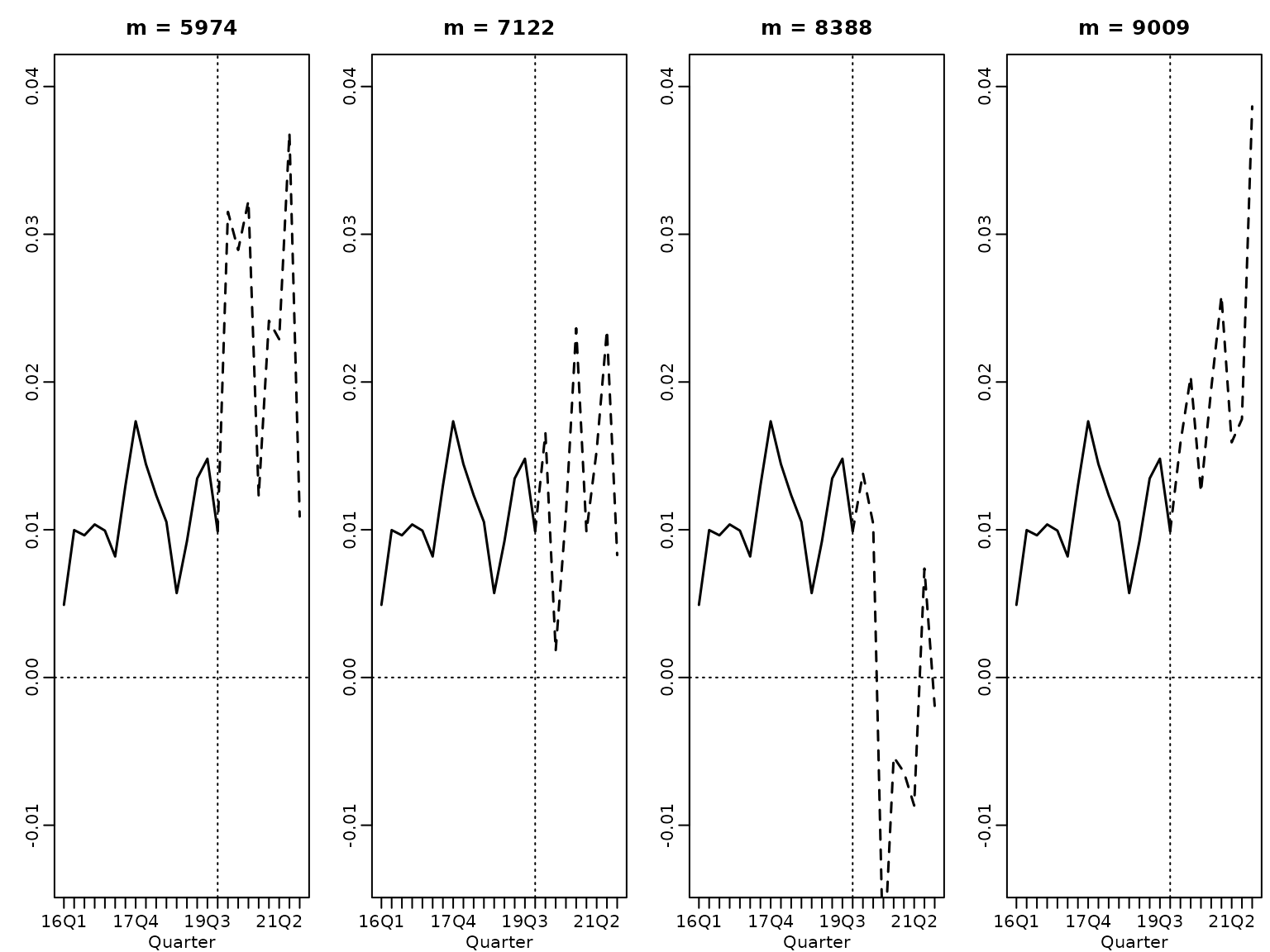
}And we visualize. First, we plot four randomly selected paths.
horizon <- 8
ats <- seq(1, 3 * horizon)
past <- as.numeric(substring(gsub("-.*", "", tail(names(y), 2 * horizon)), 3))
years <- c(past, tail(past, 1) + rep(1:(horizon / 4), each = 4))
labs <- paste0(years, "Q", 1:4)
these <- sort(sample.int(M, 4))
for (i in 1:4) {
plot(tail(y, 2 * horizon), main = paste("m =", these[i]), type = "l",
xlim = c(1, 3 * horizon), ylim = range(tail(y, 2 * horizon), yfs[these, ]),
ylab = "", xlab = "Quarter", lwd = 1.5, xaxt = "n")
axis(side = 1, at = ats, labels = labs[ats])
lines((2 * horizon):(3 * horizon), c(tail(y, 1), yfs[i, ]), lty = 2, lwd = 1.5)
abline(v = 2 * horizon, lty = 3)
abline(h = 0, lty = 3)
}
And now we plot the predictions.
par(mfrow = c(1, 1))
plot(tail(y, 2 * horizon), type = "l", xlim = c(1, 3 * horizon), lwd = 1.5,
ylim = range(quants), ylab = "U.S. GDP growth", xlab = "Quarter",
xaxt = "n", main = "Historical time series and predictive intervals")
axis(side = 1, at = ats, labels = labs[ats])
xs <- (2 * horizon + 1):(3 * horizon)
lines(xs, quants["50%", ], lwd = 1.5, col = 2, lty = 2)
pxs <- c(xs[1], xs, rev(xs))
polygon(pxs, c(quants["25%", 1], quants["75%", ], rev(quants["25%", ])),
col = rgb(1, 0, 0, .2), border = NA)
polygon(pxs, c(quants["10%", 1], quants["90%", ], rev(quants["10%", ])),
col = rgb(1, 0, 0, .2), border = NA)
polygon(pxs, c(quants["5%", 1], quants["95%", ], rev(quants["5%", ])),
col = rgb(1, 0, 0, .2), border = NA)
abline(h = 0, lty = 3)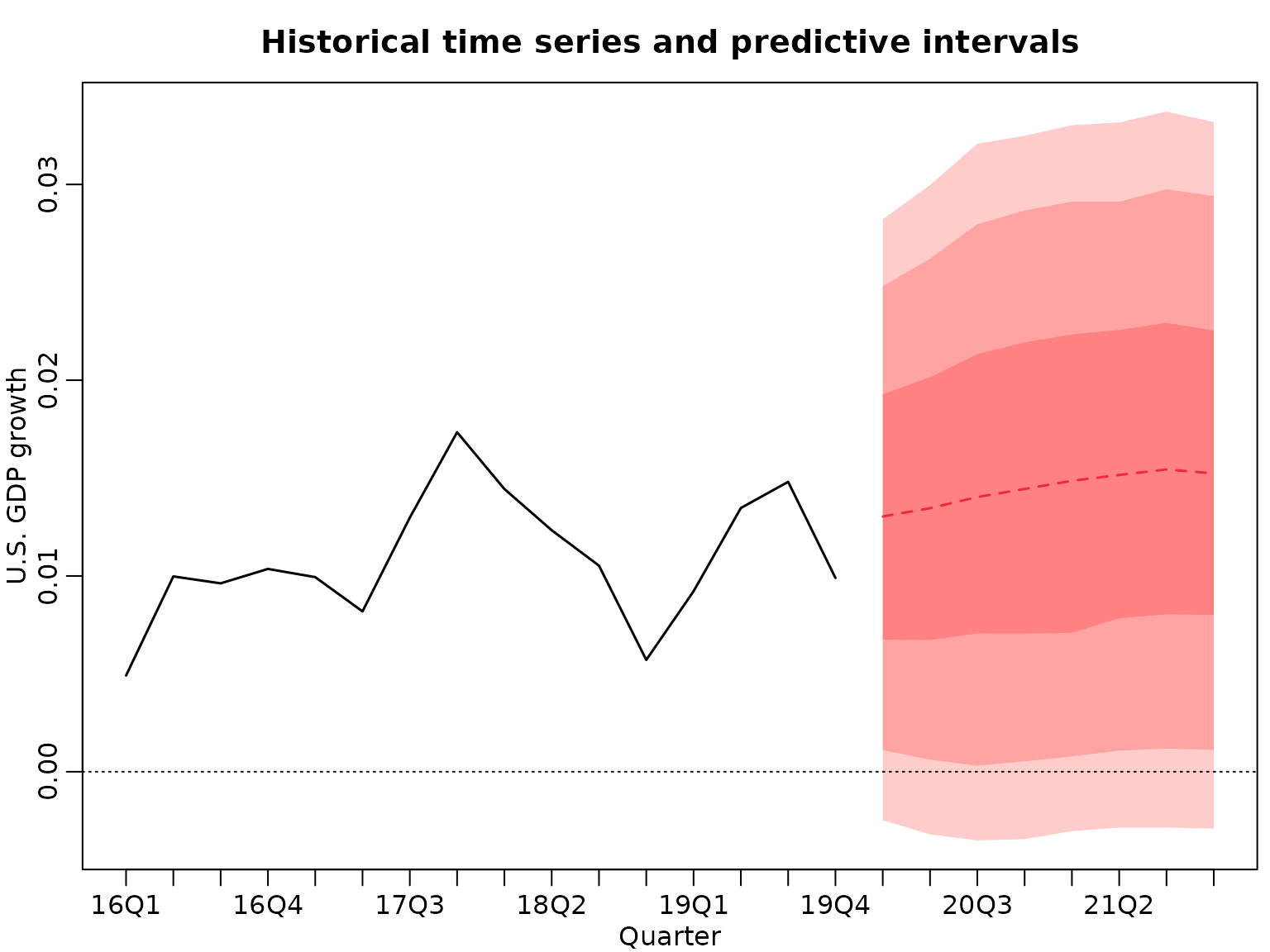
Let us compute the probability of seeing negative growth rates a least once in eight quarters. Not that this is highly nonlinear.
Another example of a nonlinear function of the predicted log returns is the level, which, given our simulation-based approach, is easy to compute.
We visualize.
par(mfrow = c(1, 1))
plot(tail(gdp, 2 * horizon), type = "l", xlim = c(1, 3 * horizon),
ylim = range(quants, tail(gdp, 2 * horizon)), ylab = "U.S. GDP",
xlab = "Quarter", lwd = 1.5, xaxt = "n",
main = "Historical time series and predictive intervals")
axis(side = 1, at = ats, labels = labs[ats])
xs <- (2 * horizon + 1):(3 * horizon)
lines(xs, quants["50%", ], lwd = 1.5, col = 2, lty = 2)
pxs <- c(xs[1], xs, rev(xs))
polygon(pxs, c(quants["25%", 1], quants["75%", ], rev(quants["25%", ])),
col = rgb(1, 0, 0, .2), border = NA)
polygon(pxs, c(quants["10%", 1], quants["90%", ], rev(quants["10%", ])),
col = rgb(1, 0, 0, .2), border = NA)
polygon(pxs, c(quants["5%", 1], quants["95%", ], rev(quants["5%", ])),
col = rgb(1, 0, 0, .2), border = NA)
abline(h = 0, lty = 3)